Simultaneous Detection of Feces-specific Bacteriophages of Bacteroides fragilis with a Duplex PCR Assay
Main Article Content
Abstract
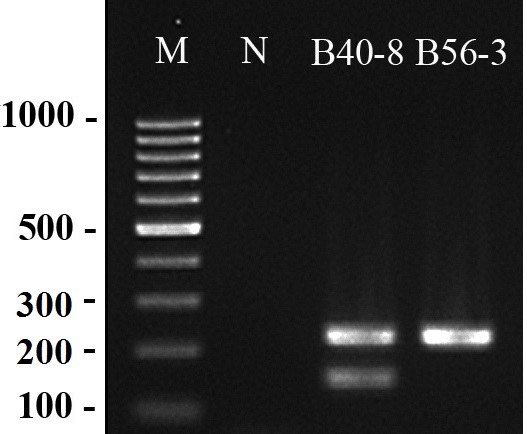
Bacteriophages of the Bacteroides fragilis strains HSP40 and RYC2056 are used as indicators of human-specific and general (non-host specific) fecal pollution in water bodies. However, conventional anaerobic cultivation methods require 1-2 days of incubation. To overcome this limitation, in this study, we developed a DNA-based method to simultaneously detect representative bacteriophages (B40-8 and B56-3) that infect B. fragilis strains HSP40 and RYC2056, respectively. Both phages yielded a 224-bp amplicon with the primer pair BT5414/BT5415, and an additional 152-bp PCR product was observed for B40-8 with the primer pair BT5579/BT5580. The detection limits for B40-8 and B56-3 were 10-5 and 10-4 ng of pure DNA, and 1 and 50 ng of DNA template when 5 and 5,000 PFU/mL were spiked into distilled water, respectively. The assay exhibited a higher sensitivity for sewage samples, with < 0.1 and 15 PFU/mL of phages infecting HSP40 and RYC2056, respectively. The assay did not produce false positive results for the Bacteroides phages PG76, HB13, and GA17 or for the enterococcal phages AIM06 and SR14. The assay also detected RYC2056 phages that were isolated from sewage samples and the phage B40-8 when it was spiked into raw sewage. Thus, the newly developed PCR assay demonstrated potential for the environmental monitoring of Bacteroides bacteriophages, decreasing the analysis time to a few hours.
Article Details
Published articles are under the copyright of the Environment and Natural Resources Journal effective when the article is accepted for publication thus granting Environment and Natural Resources Journal all rights for the work so that both parties may be protected from the consequences of unauthorized use. Partially or totally publication of an article elsewhere is possible only after the consent from the editors.